Part 1: Template README and Reproducible Practices
The Plan
| Time | April 11, 2026 (Saturday) |
|---|---|
| 8:00 | Breakfast |
| 9:00 | Introduction (with World Bank intro) |
| 10:00 | Reproducible Practices, Template README |
| 11:00 | Data provenance, Data Citations |
| 12:00 | Lunch Break |
| 13:00 | What will you be doing in the Lab |
| 14:00 | Command Line/Git/Markdown/Version control |
| 15:00 | A prototypical replication report |
| 16:00 | A walkthrough of the workflow |
| 17:00 | How to run Stata code |
| 18:00 | Restaurant |
Overview
Part 1:
- Reproducible practices
- The role of the template README
- Ideal directory and data structure
- Some programming practices
Reproducible practices
Version code and results
While we are assessing reproducibility of others, our work must be reproducible as well.
- Use versioning. We will use
git. We will show you how. - Keep a log of your work
- Git will log some things
- You will keep a high-level log (notes) of your work in the report (
REPLICATION.md)
- Commit often (at least once a day)
Part 1 | Ideal structure
What do we tell researchers?
Here’s a few generic guidelines for researchers. You will be on the lookout for these things!
Generic project setup
Basic project setup
Structure your project
- Data inputs
- Data outputs
- Code
- Paper/text/etc.
Version your project (git)!
Track metadata
- cite articles you reference
- cite data sources you use
- We will get back to data citations later!
Project setup examples
/inputs
/outputs
/code
/paper/datos/
/brutos
/limpiados
/finales
/codigo
/articuloIt doesn’t really matter, as long as it is logical. We will get to how this translates to confidential or big data in a moment!
Computational Empathy
Consider how the next person will (be able to) compute
- You don’t know who that is
- You don’t know what they don’t know
- Will not have any of your add-on packages/ libraries/ etc. pre-installed
- Don’t force them to do tedious things
It might be “Future You!”
It IS you
The replicator is the first (?) reader of the instructions who will need to reproduce the analysis.
Streamlining
- Master script preferred
- Least amount of manual effort
- No manual copying of results
- (dynamic documents!)
- Write out/save tables and figures using packages
- Clear instructions
- No manual install of packages
- Use a script to create all directories, install all necessary packages/requirements/etc.
Reproducibility
- No manual manipulation
- “Change the parameter to 0.2, then run the code again”
- Use functions, ado files, programs, macros, subroutines
- Use loops, parameters, parameter files to call those subroutines
- Use placeholders (globals, macros, libnames, etc.) for common locations (
$CONFDATA, $TABLES, $CODE)
- Compute all numbers in package
- No manual calculation of numbers
- Use cross-platform programming practices
Cross-platform programming practices 1
Use programming-language specific code as much as possible
Avoid
or
Cross-platform programming practices 1
Most languages have appropriate code:
R:
Stata:
Cross-platform programming practices 2
Use neutral pathnames (mostly forward slashes)
R: Use functions to combine paths (and/or use forward slashes), packages to make code more portable.
basepath <- rprojroot::find_root(rprojroot::has_file("README.md"))
data <- read.dta(file.path(basepath,"path","data.dta"))Stata: always use forward slashes, even on Windows
global rootdir "/my/computer"
use "$rootdir/path/data.dta"Cross-platform programming practices : better!
Use neutral pathnames (mostly forward slashes)
R: Use functions to combine paths (and/or use forward slashes), packages to make code more portable.
basepath <- rprojroot::find_root(rprojroot::has_file("README.md"))
data <- read.dta(file.path(basepath,"path","data.dta"))Stata: always use forward slashes, even on Windows
global rootdir : pwd
use "$rootdir/path/data.dta"More complex Data structures
Back to the TIER protocol
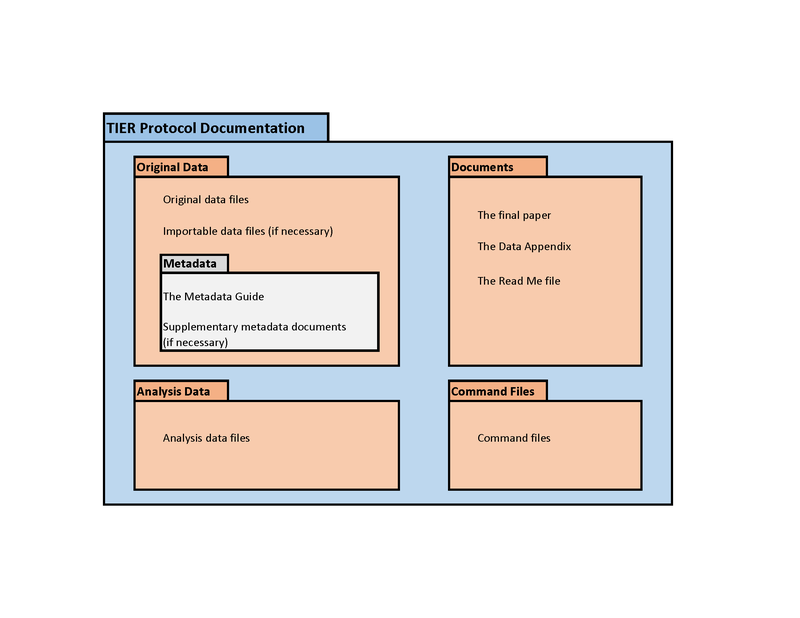
TIER Protocol again
Generic suggested data structure
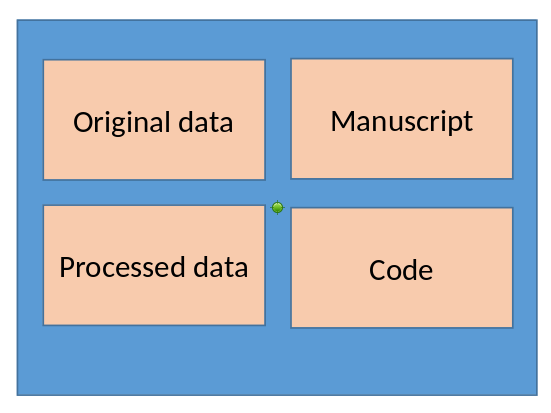
Simplified replication package structure
When data are big/in the cloud

Data are big
When data are confidential

Confidential data
When data are confidential

Confidential data in enclaves
Project setup examples
This may no longer work:
/data/
/raw
/clean
/final
/code
/articleBut this might
/project123/
/data/
/raw
/clean
/final
/code
/article
/confidential (read-only)
/taxes (read-only)
/wages (read-only)Stata configuration files
File structure thus becomes more complex, but fundamentally not so different:
global taxdata "/confidential/taxes"
global salarydata "/confidential/wages"
global outputdata "/project/data/clean" // this is where you would write the data you create in this project
global results "/project/article" // All tables for inclusion in your paper go here
global programs "/project/code" // All programs (which you might "include") are to be found hereStata configuration files
Or even more robust:
global rootdir "/project123"
global confbase "/data/provided"
global project "$rootdir/project"
global taxdata "$confbase/taxes"
global salarydata "$confbase/wages"
global outputdata "$project/data/clean" // this is where you would write the data you create in this project
global results "$project/article" // All tables for inclusion in your paper go here
global programs "$project/code" // All programs (which you might "include") are to be found hereNext up: README